Applied science
Applying powerful AI to solve grand challenges in natural science for the benefit of the world.

Applying powerful AI to solve grand challenges in natural science for the benefit of the world.
About the team
Computer science and natural science are complementary: breakthroughs in one drive remarkable advances in the other. Google’s Applied Science organization aims to cross-fertilize these two fields across a wide range of scientific disciplines.
Within Applied Science, the Climate & Energy team focuses on technology to mitigate climate change, while the Science AI team leverages AI to accelerate progress in natural sciences. By tackling grand challenges from reconstructing the brain to inventing zero-carbon energy sources, Applied Science advances both computer science and scientific research simultaneously, including using AI to automate code generation for computational experiments.
Team focus summaries
The Climate & Energy team deploys AI and Google's extensive computing power to combat climate change. We aim to provide the scientific insights and technical tools necessary for global climate mitigation and adaptation. We also investigate both technological and nature-based methods for atmospheric CO2 removal, including sequestration in the oceans.
Primary research goals include:
- Accelerating clean energy innovations, such as nuclear fusion.
- Developing strategies to mitigate various warming agents, including aviation-related contrails.
- Employing AI alongside remote sensing to monitor and manage climate-related features like wildfires, methane leaks, and water resources.
Google researchers and their collaborators use computational power to drive breakthroughs in climate science, environmental modeling and biodiversity mapping. Key initiatives include:
- NeuralGCM: Combining fundamental physics laws with an ML model trained on historical weather and satellite data to create a hybrid atmospheric model. This open-source model, NeuralGCM, is fast, efficient and can reproduce real-world weather events with high accuracy.
- Targeting climate forecasting: The development of smaller-scale, specialized models for particular weather and climate prediction needs.
- Global biodiversity mapping: Using LLMs and geospatial models to map the habitats of hundreds of thousands of species to facilitate local, national, and international conservation efforts.
For more than a decade, the genomics group at Google has leveraged AI to enable more accurate analyses of genetic data. As genetic sequencing technology has evolved, so have our analysis tools. Our suite of open-source AI models such as DeepVariant, DeepConsensus, DeepPolisher and, most recently, DeepSomatic, support genetic analysis in research, medicine and conservation.
Today we also have single-cell sequencing to detect when genes are active, as well as data from medical imaging, fitness trackers, patient history and more. The multimodal biology group at Google seeks ways to combine these various data sources to unlock medical insights.
The human brain contains billions of neurons and fully mapping it remains out of reach. However, Google Research is precisely mapping the connections between every cell in simpler brains to someday understand and even treat mental illnesses. In collaboration with external research partners, we develop AI-based methods to efficiently reconstruct neurons from 2D images and create tools, such as Neuroglancer, to view and analyze that data.
We are working with our research partners to create increasingly ambitious brain maps for model organisms such as the fruit fly, zebrafish, the mouse and even sections of the human brain. Research groups are now using those maps to predict the neural activity and behavior of model organisms.
We develop foundational tools for scientific discovery that leverage AI-enabled coding and LLMs to democratize access to advanced analysis and automate complex research tasks.
Featured publications
Highlighted work
-
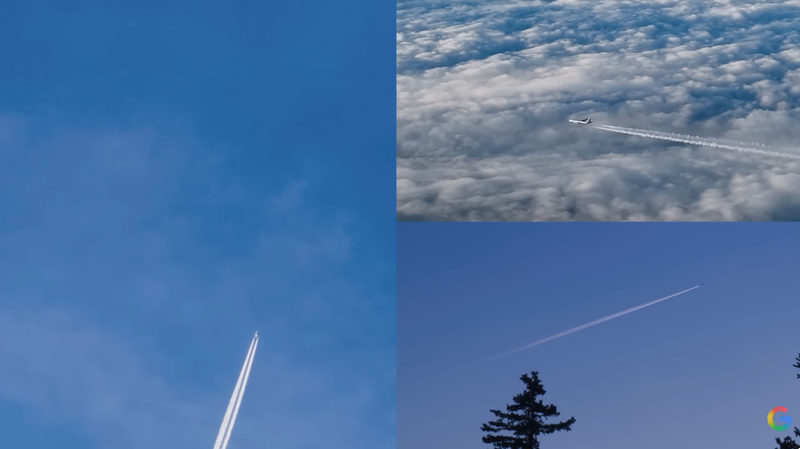 Project ContrailsA cost-effective and scalable approach to mitigate aviation’s climate impact.
Project ContrailsA cost-effective and scalable approach to mitigate aviation’s climate impact. -
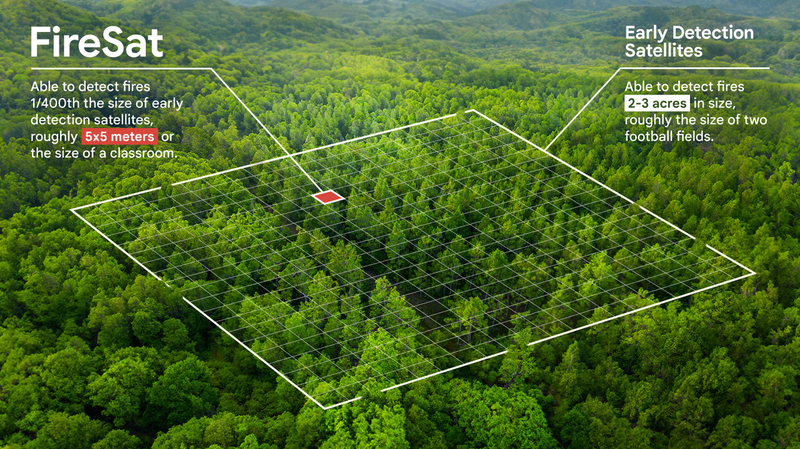 FireSAT: Wildfire DetectionUsing high-res multispectral satellite imagery and AI to provide near real-time insights on wildfires.
FireSAT: Wildfire DetectionUsing high-res multispectral satellite imagery and AI to provide near real-time insights on wildfires. -
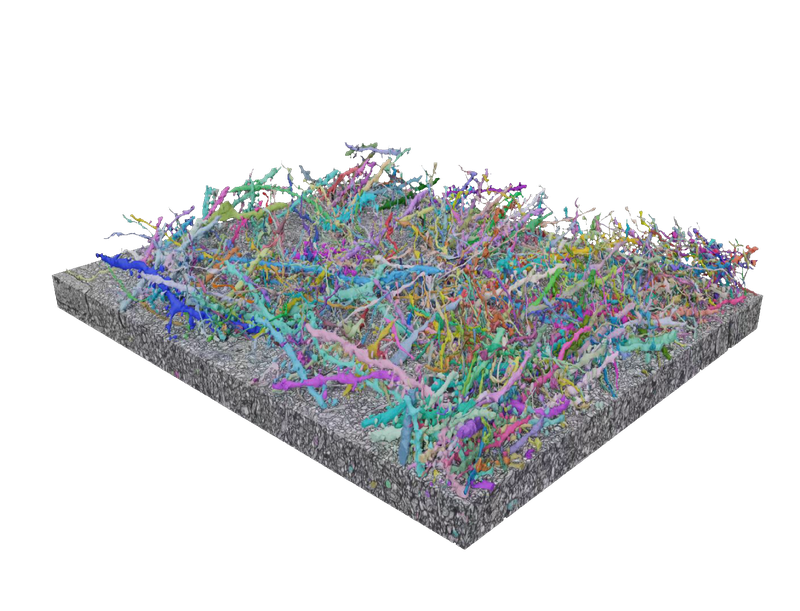 Neuroscience and ConnectomicsDriving progress toward precisely mapping the connection between every cell in the brain.
Neuroscience and ConnectomicsDriving progress toward precisely mapping the connection between every cell in the brain. -
 Models of the WorldA hybrid model of the atmosphere that combines physics and AI to deliver fast, accurate weather and climate simulations.
Models of the WorldA hybrid model of the atmosphere that combines physics and AI to deliver fast, accurate weather and climate simulations. -
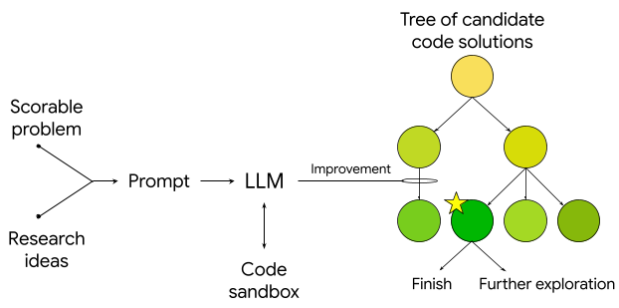 Scientific ToolsAccelerating scientific discovery with AI-powered empirical software
Scientific ToolsAccelerating scientific discovery with AI-powered empirical software
Some of our locations
Some of our people
-

John C. Platt
- General Science
- Machine Intelligence
-

Michael P Brenner
- General Science
- Machine Intelligence
- Health & Bioscience
-

Christopher H Van Arsdale
- Economics and Electronic Commerce
- General Science
- Distributed Systems and Parallel Computing
-

Subhashini Venugopalan
- General Science
- Machine Intelligence
- Machine Perception
-

Nita Goyal
- General Science
- Machine Intelligence
-

Mike Tyka
- Human-Computer Interaction and Visualization
- Machine Intelligence
- Machine Perception
-

Cory McLean
- General Science
- Machine Intelligence
- Health & Bioscience
-

Kevin McCloskey
- General Science
- Machine Intelligence
-

Viren Jain
- General Science
- Machine Perception




